Introduction
Materials and Methods
Experimental materials
Ethyl acetate fractionation
Cell culture
Immunoblotting
Revers transcription-polymerase chain reaction (RT-PCR)
Quantitative PCR Analysis
Statistical analysis
Results and Discussion
Introduction
The skin consists of the dermis and epidermis. The epidermis protects against various physical, chemical and mechanical stimuli, and excessive divergence of moisture (Chung et al., 2003; Fisher et al., 2002; Kim et al., 2014). Melanin is an important substance produced in the melanocytes located in the basal layer of the skin epidermis. After melanin is produced, it affects the formation of skin and hair pigments (Ali, 2010). It particularly protects the skin from ultraviolet (UV) damage, acting as a natural antioxidizing, toxic elimination, and powerful anion chelating agent (Regnault et al., 2015). Abnormal melanogenesis, which causes the disease, vitiligo or hyperpigmentation, is known to be caused by intrinsic factors, such as genetic predisposition, hormonal changes (such as change in estrogen levels), liver disease, tumor, and cancer, and extrinsic factors, such as excessive exposure to UV and toxic substances, which majorly result in hyperpigmentation (Ali, 2010). Abnormal formation of the skin pigment is caused by oxidative stress due to excessive active oxygen species generated by UV rays, and the antioxidants that eliminate them have also been reported to be effective in inhibiting melanin formation (Eberlein-Konig et al., 1998). Tyrosinase is an enzyme that regulates both the stimulation and inhibition of melanin production. Its expression is regulated by microphthalmia-associated transcription factor (MITF), which regulates the differentiation, proliferation and growth of melanocytes (Busca and Ballotti, 2000; Garraway et al., 2005). MITF regulation is initiated by signaling after α-melanocyte-stimulating hormone (α-MSH) binds to the Melanocortin 1 receptor (MC1R). The MC1R promotes the signaling of mitogen-activated protein kinase (MAPK), including extracellular signaling-regulated kinase (ERK) and p38. Melanin synthesis is also regulated by Phosphatidylinositol-3-kinase (PI3K / AKT) signaling (Onkoksoong et al., 2018; Zhou et al., 2017).
Pinus densiflora is an evergreen conifer, native to the Far East, including Korea, Japan, and China. These countries use pine leaves, pollen and bark for medicinal or edible purposes (Lee, 2003). P. densiflora contains terpenes, such as α-pinene, β-pinene, and camphene, in addition to polyphenols, flavonoids, inorganic and organic components, and vitamins (Chung et al., 1996). There are many reports on the functional effects of P. densiflora leaf extracts, including bioactivity, antioxidation, immunity, pathogenicity, antimicrobial activity against food-borne microorganisms, food efficacy, skin whitening, and anti-aging (Choi et al., 1997; Sung and Kim, 2005; Yoo et al., 2004). However, the inhibitory effect of the pine cone on melanin synthesis has not been reported. In this study, we investigated the inhibitory effect of PEF on melanin formation.
Materials and Methods
Experimental materials
Pine cone of Pinus densiflora (voucher number: CLP09- 27) was collected from Cheollipo arboretum, 187, Cheollipo 1-gil, Sowon-myeon, Taean-gun, Chungcheongnam-do, Republic of Korea. Acetonitrile, chloroform, dimethyl sulfoxide (DMSO), ethyl acetate, methanol, and petroleum ether (HPLC- grade) were purchased from Merck (Darmstadt, Germany). Dulbecco’s modified Eagle’s medium (DMEM), 10% (v/v) fetal bovine serum, penicillin/streptomycin, and trypsin were purchased from Hyclone (Logan, UT, USA). The primary and secondary antibodies were purchased from Abcam (Cambridge, UK) and Santa Cruz Biotechnology (Dallas, TX, USA). All electrophoresis related chemicals were purchased from Bio-Rad Labs (Hercules, CA, USA). All other chemicals were purchased from Sigma-Aldrich (St. Louis, MO, USA), unless otherwise specified. Standard acteoside was purchased from Sigma- Aldrich.
Ethyl acetate fractionation
Pine cone (50 g) was extracted with 80% methanol (1 L) and the extract was concentrated by using a vacuum evaporator (N-1110S, EYELA, Shanghai, China). The aqueous residue was fractionated with petroleum ether, and ethyl acetate. The ethyl acetate fraction (0.13 g) was concentrated, freeze dried and stored in a refrigerator until use at 4℃. PEF was dissolved in DMSO and treated to cells. DMSO was used as a vehicle and the final DMSO concentration did not exceed 0.1% (v/v).
Cell culture
B16F10 cells were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA) and cultured in DMEM supplemented with 10% (v/v) fetal bovine serum, 100 U/mL penicillin, and 100 ㎍/mL streptomycin. The cells were maintained at 37℃ under a humidified atmosphere containing 5% CO2.
Immunoblotting
B16F10 cells were cultured in 6-well plates at 37℃ in an incubator with a humidified atmosphere containing 5% CO2. The cells were washed with 1 × phosphate-buffered saline, lysed in radio immuno precipitation assay buffer (Thermo Scientific, Waltham, MA, USA) supplemented with protease inhibitor cocktail (Sigma-Aldrich), and subsequently centrifuged at 12,000 × g for 15 min at 4℃. The protein concentration of the sample was determined by the Bradford protein assay (Bio-Rad, Hercules, CA, USA). The proteins were mixed with Laemmli buffer and boiled at 95℃ for 5 min. The proteins were then separated using 10% SDS-PAGE and transferred to a polyvinylidene fluoride membrane (Bio- Rad). The membranes were blocked with 5% nonfat dry milk in Tris-buffered saline containing 1% Tween 20 (TBS-T) for 30 min and then incubated with specific primary antibodies in 3% nonfat dry milk at 4℃ overnight. Subsequently the blots were washed three times with TBS-T and the blots were incubated with horseradish peroxidase-conjugated immunoglobulin G for 1 h. The chemiluminescence was detected using ECL Western blotting substrate (Bio-Rad) and visualized with Chemi-Doc (Bio-rad, Hercules, CA, USA).
Revers transcription-polymerase chain reaction (RT-PCR)
The total RNA was extracted from B16F10 cells using NucleoSpin® RNA Plus (Macherey-Nagel, Düren, Germany) and was synthesized using ReverTra Ace –α- (Toyobo, Osaka, Japan) according to the manufacturer’s protocol for cDNA synthesis. PCR was carried out using Quick Taq® HS DyeMix (Toyobo).
Quantitative PCR Analysis
The total RNA was extracted using RNeasy® Plus Mini kit (Qiagen, Hilden, Germany) and quantified using Quanti Fluor® RNA system (Promega, Madison, USA) and Quantus™ Fluorometer (Promega). The RNA was synthesized using ReverTra Ace –α- (Toyobo, Osaka, Japan) according to the manufacturer’s protocol for cDNA synthesis. The cDNAs were then used as templates for Quantitative polymerase chain reaction (qPCR) using QuantiTect® SYBR Green PCR Kit (Qiagen). following the manufacturer’s instructions. To quantify the mRNA expression, quantitative polymerase chain reaction (qPCR) analysis was performed with a 7500 Real-Time PCR system (Applied Biosystems, CA, USA). qPCR primers were designed using the Primer3 program (Table 1). Data were analyzed using 7500 Software 2.3 (Applied Biosystems).
Table 1. Primer sequence for PCR
Statistical analysis
Statistical analysis was carried out using SPSS 18.0 (SPSS, Inc., Chicago, IL, USA). All experiments were performed at least three times. The data obtained from experiments carried out in triplicates are shown as the mean ± SD. Differences were considered statistically significant when the P value was < 0.05.
Results and Discussion
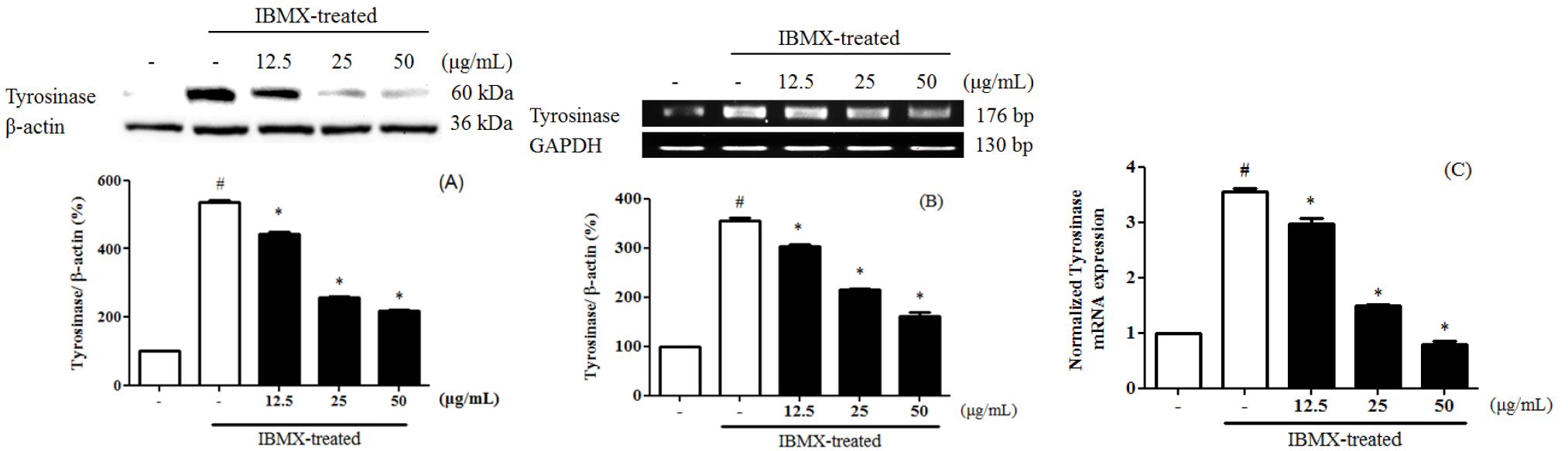
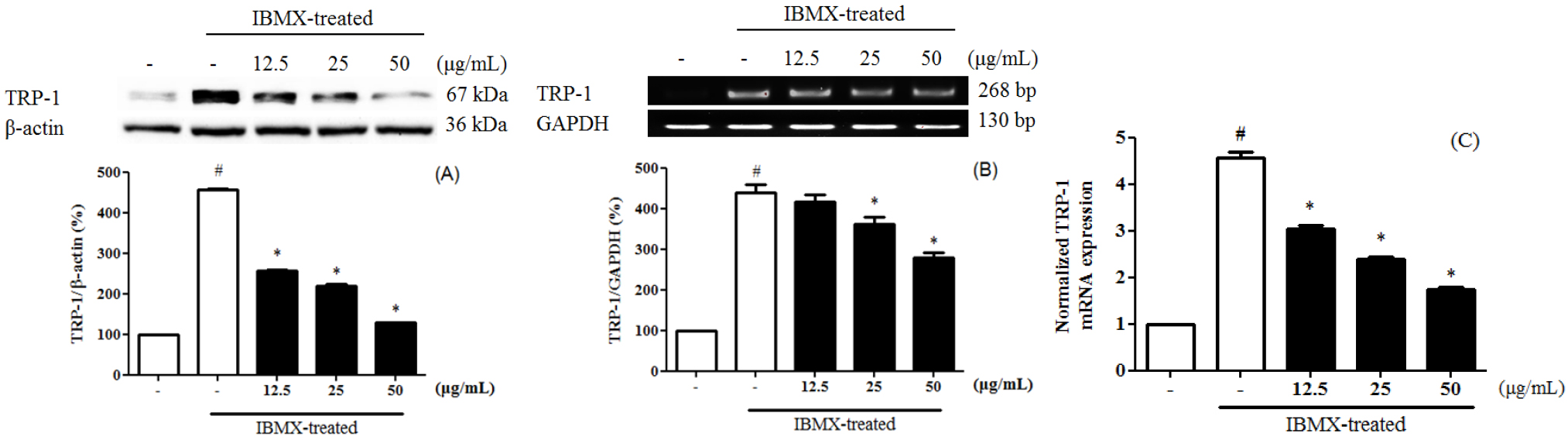
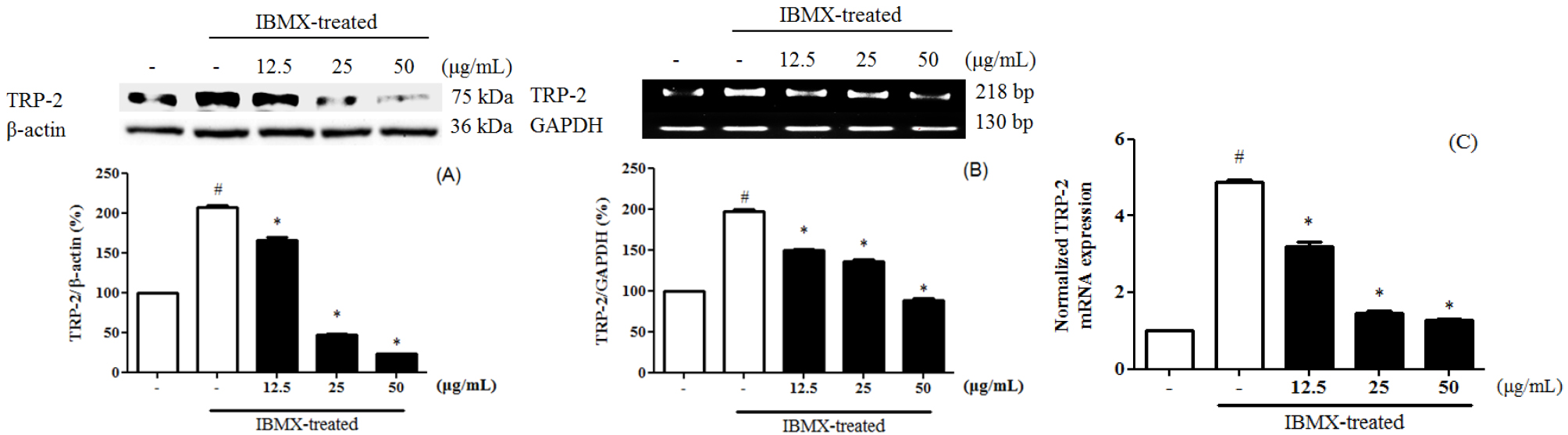
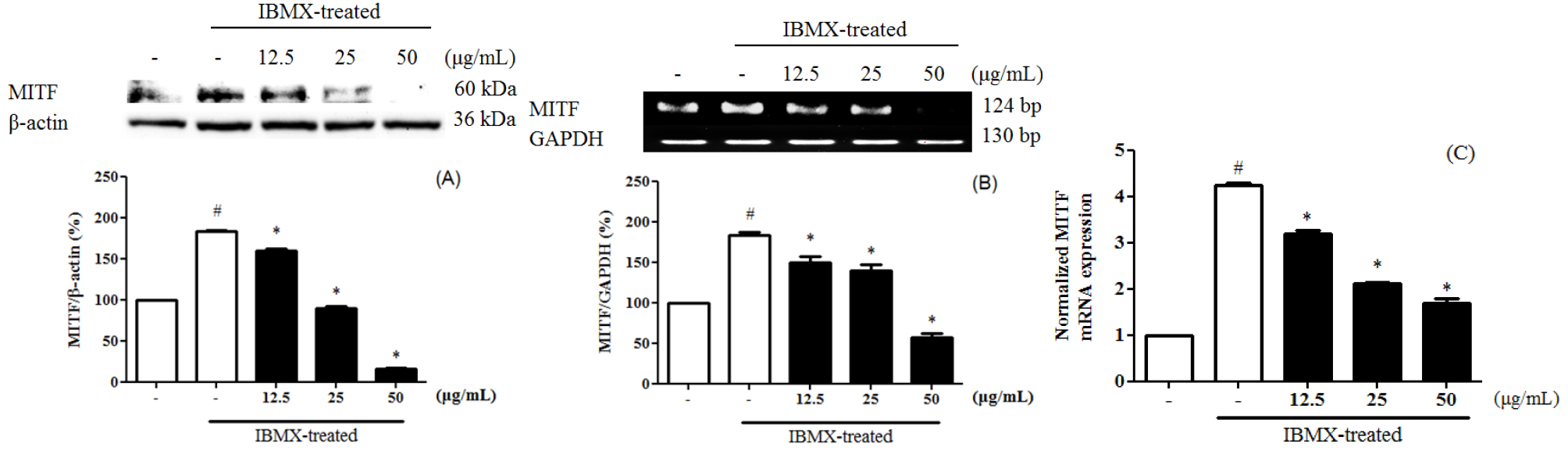
Melanin is divided into eumelanin and pheomelanin during the synthesis process. Inhibition of tyrosinase activity is important for melanogenesis and synthesis, since melanin biosynthesis is known to be an early reaction (Hearing and Tsukamoto, 1991), which is known to convert tyrosine to L-3,4-dihydroxyphenylalanine (DOPA) by binding to the hydroxyl group of the substrate tyrosine (Jimenez-Cervantes et al., 1994). 3-Isobutyl-1-methylxanthine (IBMX) elevated the level of intracellular cyclic-AMP (cAMP), a key messenger in the regulation of skin pigmentation. Treatment of IBMX in B16F10 stimulated cAMP-mediated melanin production. (Busca and Ballotti, 2000; Im et al., 1998). To evaluate the tyrosinase expression of PEF in IBMX-induced B16F10 cells, B16F10 cells were pre-treated with PEF and IBMX. Tyrosinase expression in IBMX-induced B16F10 increased compared to the untreated group. B16F10 cells were treated with PEF (12.5, 25, and 50 ㎍/mL) for 24 hours. Compared with the PEF-untreated group, PEF inhibited tyrosinase expression in a concentration-manner, confirmed by immunoblotting (Fig. 1A), RT-PCR analysis (Fig. 1B) and qPCR analysis (Fig. 1C). Compared to the basal level, IBMX treatment significantly increased tyrosinase expression by 535.8 ± 5.6%. However, PEF inhibited tyrosinase expression at 216.9 ± 4.8% at 50 ㎍/mL. Compared to the basal level, IBMX treatment significantly increased tyrosinase mRNA levels by 221.3 ± 6.5%. PEF inhibited tyrosinase mRNA levels at 167.1 ± 4.5% at 50 ㎍/mL. IBMX-induced B16F10 cells were up-regulated the tyrosinase mRNA levels by 3.55 ± 0.06-fold, compared to the untreated group. The tyrosinase mRNA levels were down-regulated by PEF treatments by 0.8 ± 0.05-fold at 50 ㎍/mL. Also, melanogenesis is regulated by three specific enzymes: tyrosinase, tyrosinase-related protein-1 (TRP-1), and tyrosinase-related protein-2 (TRP-2). During melanogenesis, TRP-1 and TRP-2 have functional roles. TRP-1 catalyzes the oxidation of 5,6-dihydroxyindole-2- carboxylic acid (DHICA), and TRP-2 (dopachrome tautomerase) catalyzes the conversion of dopachrome to DHICA (Sato et al., 2008). In addition, the two enzymes are regulated by a specific transcription factor, microphthalmia-associated transcription factor (MITF) (Hasegawa et al., 2010; Widlund and Fisher, 2003). To evaluate the TRP-1 expression of PEF in IBMX-induced B16F10 cells, B16F10 cells were pre- treated with PEF and IBMX. TRP-1 expression in IBMX- induced B16F10 increased compared to the untreated group. Compared with the PEF-untreated group, PEF inhibited TRP-1 expression in a concentration-manner, confirmed by immunoblotting (Fig. 2A), RT-PCR analysis (Fig. 2B) and qPCR analysis (Fig. 2C). Compared to the basal level, IBMX treatment significantly increased TRP-1 expression by 456.0 ± 8.9%. However, PEF inhibited TRP-1 expression at 128.0 ± 0.2% at 50 ㎍/mL. Compared to the basal level, IBMX treatment significantly increased TRP-1 mRNA levels by 439.2 ± 19.4%. However, PEF inhibited TRP-1 mRNA levels at 278.6 ± 12.3% at 50 ㎍/mL. IBMX-induced B16F10 cells were up-regulated the TRP-1 mRNA levels by 4.56 ± 0.12- fold, compared to the untreated group. The TRP-1 mRNA levels were down- regulated by PEF treatments by 2.38 ± 0.05-fold at 50 ㎍/mL. To evaluate the TRP-2 expression of PEF in IBMX-induced B16F10 cells, B16F10 cells were pre-treated with PEF and IBMX. TRP-2 expression in IBMX- induced B16F10 increased compared to the untreated group. Compared with the PEF-untreated group, PEF inhibited TRP-2 expression in a concentration-manner, confirmed by immunoblotting (Fig. 3A), RT-PCR analysis (Fig. 3B) and qPCR analysis (Fig. 3C). Compared to the basal level, IBMX treatment significantly increased TRP-2 expression by 206.6 ± 2.7%. However, PEF inhibited TRP-2 expression at 47.3 ± 0.5%, and 22.8 ± 0.2% at 25, and 50 ㎍/mL. Compared to the basal level, IBMX treatment significantly increased TRP-2 mRNA levels by 197.6 ± 2.5%. However, PEF inhibited TRP-2 mRNA levels at 88.1 ± 2.2% at 50 ㎍/mL. IBMX- induced B16F10 cells were up-regulated the TRP-2 mRNA levels by 4.87 ± 0.08-fold, compared to the untreated group. The TRP-2 mRNA levels were down-regulated by PEF treatments by 1.27 ± 0.05-fold at 50 ㎍/mL. To evaluate the MITF expression of PEF in IBMX-induced B16F10 cells, B16F10 cells were pre-treated with PEF and IBMX. MITF expression in IBMX-induced B16F10 increased compared to the untreated group. Compared with the PEF-untreated group, PEF inhibited MITF expression in a concentration-manner, confirmed by immunoblotting (Fig. 4A), RT-PCR analysis (Fig. 4B) and qPCR analysis (Fig. 4C). Compared to the basal level, IBMX treatment significantly increased MITF expression by 182.9 ± 2.0%. However, PEF inhibited MITF expression at 16.3 ± 0.5% at 50 ㎍/mL. Compared to the basal level, IBMX treatment significantly increased MITF mRNA levels by 183.5 ± 4.0%. However, PEF inhibited MITF mRNA levels at 57.3 ± 4.6% at 50 ㎍/mL. IBMX-induced B16F10 cells were up-regulated the MITF mRNA levels by 4.24 ± 0.05-fold, compared to the untreated group. The MITF mRNA levels were down- regulated by PEF treatments by 1.68 ± 0.12-fold at 50 ㎍/mL. MITF is regulated by several signaling pathways. Mitogen-activated protein kinases (MAPKs) regulate cell differentiation, proliferation, and cellular activities. They contain ERKs, c-Jun N-terminal kinases (JNKs), and p38, and play essential roles in melanogenesis (Jiang et al., 2009; Ye et al., 2010). Melanogenesis can be induced by several extracellular stimuli, including UV light, and chemicals, such as α-MSH, and IBMX. α-MSH combined with MC1R induces the enhanced expression of MITF, dependent on increased cAMP levels, leading to increased expression of tyrosinase and other melanogenesis-related enzymes. The end result is increased melanin synthesis. According to Lee et al. (2016), the methanol extract of pine cone showed anti- oxidant and anti-melanogenic activity. In a previous study, we investigated the ethly acetate fraction of pine cone of antioxidant activity and inhibitory effect on oxidative DNA damage (Jang et al., 2016). Bioactivities of PEF were related in various phenolic compounds and their potential antioxidant activities. In the presence of cAMP stimuli like IBMX, it up-regulated the MITF expression was related in cell differentiation, proliferation, and cellular activities. Our study indicated that PEF inhibited the tyrosinase, TRP-1, and TRP-2 expression via suppressing the IBMX-induced MITF expression. In conclusion, we suggest that PEF is a potent resource for reducing melanin synthesis, and has beneficial effects in skin whitening cosmetics. In addition, further studies with the study of cell signaling pathways are needed to determine how PEF inhibited on melanogenesis.